Introduction to coordinate partitioning with smoothEMr
partition.Rmd
library(smoothEMr)The feature-partition problem
Problem statement
We observe a data matrix where each column (feature) may follow one of two different latent orderings of the samples. Let the feature index set be
We seek a partition of features into two disjoint sets such that features in are best explained by ordering system 1, and features in are best explained by ordering system 2.
Latent ordering via responsibilities
In csmoothEM, each ordering system is represented by an
responsibility matrix
where
is the (soft) membership probability of sample
in component/bin
under ordering
.
You can view
as a soft discretization of a latent 1D ordering into
bins.
Each ordering system also has csmoothEM parameters where (in the homoskedastic csmoothEM setting used here):
- are mixture weights,
- are component means for features in the corresponding subset,
- are per-feature standard deviations,
- are per-feature smoothness strengths for a RW prior along .
(Here .)
Scoring a feature against a fixed ordering
A key ingredient in both greedy partition procedures is a feature-wise alignment score: for a candidate feature , given a fixed responsibilities matrix , we compute a scalar score that measures how well feature aligns with the ordering encoded by .
The package provides two score modes through
score_mode:
-
score_mode = "none": a plug-in (ELBO) score with fixed . -
score_mode = "ml": a collapsed (marginal-like) score with per-feature optimized.
Both modes rely on the same idea: given , we can fit feature-specific (and ) in closed form (or near closed form), then evaluate a comparable scalar objective.
Simulated example
We simulate a dataset where half of the features follow one latent ordering and the other half follow a second ordering. We also permute the feature columns to make the labels nontrivial.
sim <- simulate_two_order_gp_dataset(
N = 1000,
D = 16,
t_range = c(0, 10),
range = 5,
smoothness = 2.5,
variance = 3.0,
noise_sd = 0.05,
seed = 123
)
X <- sim$X
true_group <- sim$true_group
cat("Simulated X:", nrow(X), "x", ncol(X), "\n")
#> Simulated X: 1000 x 16
print(table(true_group))
#> true_group
#> 1 2
#> 8 8Quick visualization
Instead of a heatmap, we first plot a few representative features
from each ordering. Because permute_rows_block2=TRUE
permutes the sample order for the second block, features from ordering 2
typically look misaligned when plotted against the raw sample index.
# Pick one representative feature from each true group
idx1 <- which(true_group == 1)
idx2 <- which(true_group == 2)
set.seed(1)
j1 <- sample(idx1, 1) # a feature generated by ordering 1
j2 <- sample(idx2, 1) # a feature generated by ordering 2
x1 <- as.numeric(scale(X[, j1])) # z-score for comparability
x2 <- as.numeric(scale(X[, j2]))
# Construct the two "true" orderings for rows
t1 <- sim$t1
t2 <- sim$t2
if (!is.null(sim$row_perm_block2)) {
t2 <- t2[sim$row_perm_block2]
}
# ordering indices
o1 <- order(t1)
o2 <- order(t2)
op <- par(no.readonly = TRUE)
on.exit(par(op), add = TRUE)
par(mfrow = c(2, 2), mar = c(4, 4, 2.5, 1))
# (1) feature from ordering 1 vs ordering 1
plot(t1[o1], x1[o1], type = "l", lwd = 2,
xlab = "t1 (ordering 1)", ylab = "z-scored feature value",
main = sprintf("ord1 feature (%s)\nagainst ordering 1", colnames(X)[j1]))
# (2) feature from ordering 2 vs ordering 1
plot(t1[o1], x2[o1], type = "l", lwd = 2,
xlab = "t1 (ordering 1)", ylab = "z-scored feature value",
main = sprintf("ord2 feature (%s)\nagainst ordering 1", colnames(X)[j2]))
# (3) feature from ordering 1 vs ordering 2
plot(t2[o2], x1[o2], type = "l", lwd = 2,
xlab = "t2 (ordering 2)", ylab = "z-scored feature value",
main = sprintf("ord1 feature (%s)\nagainst ordering 2", colnames(X)[j1]))
# (4) feature from ordering 2 vs ordering 2
plot(t2[o2], x2[o2], type = "l", lwd = 2,
xlab = "t2 (ordering 2)", ylab = "z-scored feature value",
main = sprintf("ord2 feature (%s)\nagainst ordering 2", colnames(X)[j2]))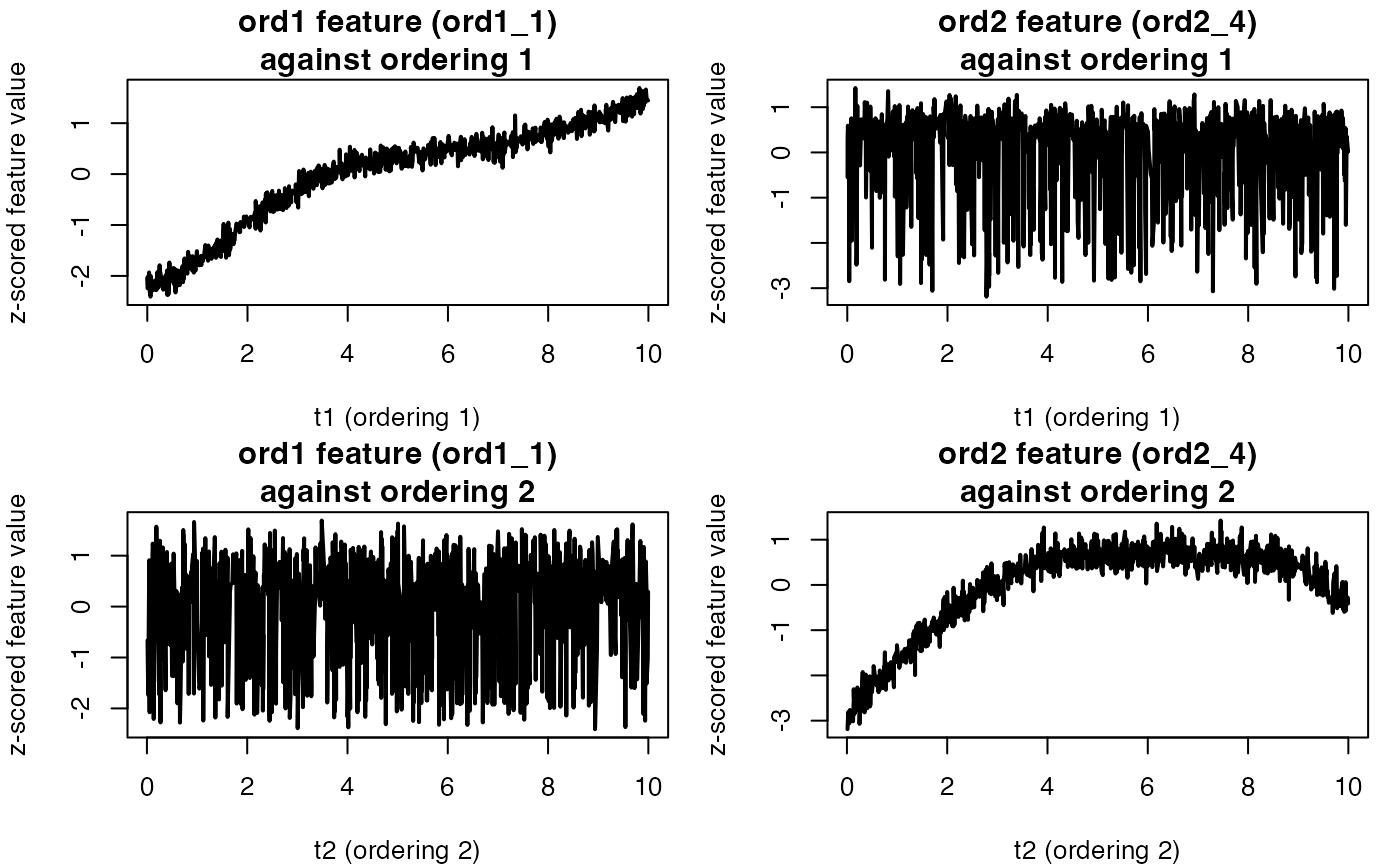
Approach 1: Forward greedy partition
Description
The forward greedy algorithm proceeds as follows:
- Choose a seed feature and fit a 1D csmoothEM ordering to obtain .
- Choose a second seed feature as the feature with the worst alignment score under , and fit a 1D csmoothEM ordering to obtain .
- While unassigned features remain:
- score each remaining feature under and ,
- pick the feature with the largest absolute score gap,
- assign it to the ordering with higher score,
- warm-start update the corresponding ordering (update and then ).
In the implementation, the ordering-specific objects are returned as
two csmooth_em fits (fit1,
fit2).
Run
set.seed(1)
res_fwd <- forward_two_ordering_partition_csmooth(
X = X,
K = 50,
score_mode = "ml",
greedy_em_refine = TRUE,
greedy_em_max_iter = 10,
verbose = 1
)
#> [Greedy-csmooth] seeds: j1=4, j2=9
#> [Greedy-csmooth] pick 3 -> ord1 (|Δ|=2357.548), remaining=13
#> [Greedy-csmooth] pick 5 -> ord1 (|Δ|=1979.785), remaining=12
#> [Greedy-csmooth] pick 13 -> ord2 (|Δ|=1898.020), remaining=11
#> [Greedy-csmooth] pick 1 -> ord1 (|Δ|=1815.358), remaining=10
#> [Greedy-csmooth] pick 6 -> ord1 (|Δ|=1547.641), remaining=9
#> [Greedy-csmooth] pick 2 -> ord1 (|Δ|=1511.510), remaining=8
#> [Greedy-csmooth] pick 7 -> ord1 (|Δ|=1344.981), remaining=7
#> [Greedy-csmooth] pick 10 -> ord2 (|Δ|=1295.184), remaining=6
#> [Greedy-csmooth] pick 16 -> ord2 (|Δ|=1062.702), remaining=5
#> [Greedy-csmooth] pick 8 -> ord1 (|Δ|=788.595), remaining=4
#> [Greedy-csmooth] pick 14 -> ord2 (|Δ|=513.117), remaining=3
#> [Greedy-csmooth] pick 11 -> ord2 (|Δ|=1374.790), remaining=2
#> [Greedy-csmooth] pick 12 -> ord2 (|Δ|=1106.892), remaining=1
#> [Greedy-csmooth] pick 15 -> ord2 (|Δ|=825.989), remaining=0
#> [Greedy-csmooth] done: |J1|=8, |J2|=8Confusion table (with label switching handled)
The partition labels 1/2 may swap. We compute both possible mappings and report the better one.
assign_raw <- as.integer(res_fwd$coord_assign)
tab1 <- table(true = true_group, est = assign_raw)
tab2 <- table(true = true_group, est = 3L - assign_raw)
acc1 <- sum(diag(tab1)) / sum(tab1)
acc2 <- sum(diag(tab2)) / sum(tab2)
if (acc2 > acc1) {
cat("Forward: using swapped labels\n")
print(tab2)
cat("Accuracy:", acc2, "\n")
} else {
cat("Forward: using original labels\n")
print(tab1)
cat("Accuracy:", acc1, "\n")
}
#> Forward: using original labels
#> est
#> true 1 2
#> 1 8 0
#> 2 0 8
#> Accuracy: 1You can also inspect the two ordering-specific fitted objects:
print(res_fwd$fit1)
#> Fitted csmoothEM object with RW(2) prior along K = 50
#> -----
#> n = 1000, d = 8, modelName = homoskedastic
#> iter = 0; init_method = forward_greedy; adaptive = ml;
#> lambda_vec: range=[1, 1], mean=1, relative = TRUE
#> ELBO last = NA; penLogLik last = NA; ML/C last = NA
print(res_fwd$fit2)
#> Fitted csmoothEM object with RW(2) prior along K = 50
#> -----
#> n = 1000, d = 8, modelName = homoskedastic
#> iter = 0; init_method = forward_greedy; adaptive = ml;
#> lambda_vec: range=[1, 1], mean=1, relative = TRUE
#> ELBO last = NA; penLogLik last = NA; ML/C last = NAApproach 2: Backward greedy partition
Description
The backward greedy algorithm starts from a single ordering fit and then moves features to a second ordering if doing so improves the alignment score.
- Fit ordering 1 on all features to obtain .
- Initialize ordering 2 from a seed feature (by default, the worst-aligned feature under ordering 1), yielding .
- Repeat:
- for each feature currently in ordering 1, compute the gain
- move the best feature if the gain is positive,
- warm-start update both orderings (update and then ).
Importantly, the implementation maintains warm-start state and does not re-run global ordering initializers after the initial construction.
Run
set.seed(1)
res_bwd <- backward_two_ordering_partition_csmooth(
X = X,
K = 50,
score_mode = "ml",
warm_iter_init = 10,
warm_iter_refit = 10,
verbose = TRUE
)
#> [backward-init] seedB=9 (ord2_1), |A|=15 |B|=1
#> [backward 01] move 13 (ord2_5) gain=1011.576 |A|=14 |B|=2
#> [backward 02] move 10 (ord2_2) gain=542.226 |A|=13 |B|=3
#> [backward 03] move 16 (ord2_8) gain=378.083 |A|=12 |B|=4
#> [backward 04] no positive gain; stopping.Confusion table (with label switching handled)
assign_raw <- ifelse(res_bwd$assign == "A", 1L, 2L)
tab1 <- table(true = true_group, est = assign_raw)
tab2 <- table(true = true_group, est = 3L - assign_raw)
acc1 <- sum(diag(tab1)) / sum(tab1)
acc2 <- sum(diag(tab2)) / sum(tab2)
if (acc2 > acc1) {
cat("Backward: using swapped labels\n")
print(tab2)
cat("Accuracy:", acc2, "\n")
} else {
cat("Backward: using original labels\n")
print(tab1)
cat("Accuracy:", acc1, "\n")
}
#> Backward: using original labels
#> est
#> true 1 2
#> 1 8 0
#> 2 4 4
#> Accuracy: 0.75